BiRAGAS Platform
Biological Retrieval-Augmented Generation Assessment System
AI-powered causal intelligence for biomarker discovery
BiRAGAS Platform
Biological Retrieval-Augmented Generation Assessment System
AI-powered causal intelligence for biomarker discovery
BiRAGAS is a scientist-guided AI platform developed by Ayass Bioscience, LLC that uncovers molecule-to-molecule causal relationships across integrated genomic, transcriptomic, and perturbation data.
By combining advanced computational modeling with expert biological interpretation, BiRAGAS provides a new standard of mechanistic evidence to support biomarker discovery and therapeutic development.
From Correlation to Causation
Most biomedical research platforms detect associations, not causal relationships. They can show that Gene A and Gene B change together, but cannot determine which gene drives the change — or whether either is truly responsible for disease.
This correlation trap limits biological insight and contributes to misdirected research and high clinical trial failure rates.
BiRAGAS addresses the core question biology has struggled to answer at scale:
Does X cause Y — or are they simply correlated?
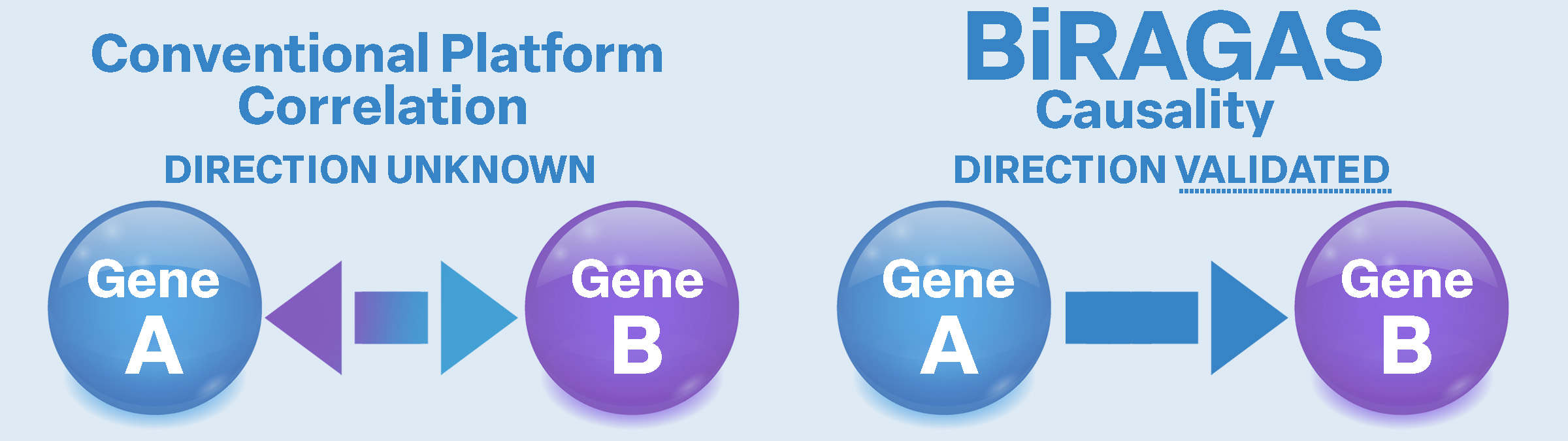
BiRAGAS infers causal relationships directly between molecular entities, rather than relying on disease labels or cohort-level associations. Disease labels are treated as cohort identifiers, not components of the model, ensuring that discoveries are grounded in underlying biological mechanisms rather than statistical correlations.
This approach enables mechanistic insights that are both biologically meaningful and clinically generalisable.

Why this matters
Clinical trial failures are often driven by incorrect target selection.
Association-based models frequently identify biomarkers rather than true disease drivers.
Regulatory approval requires mechanistic evidence.
The FDA increasingly demands causal biological understanding to support therapeutic claims.
Every Causal Claim Validated Across Independent Streams
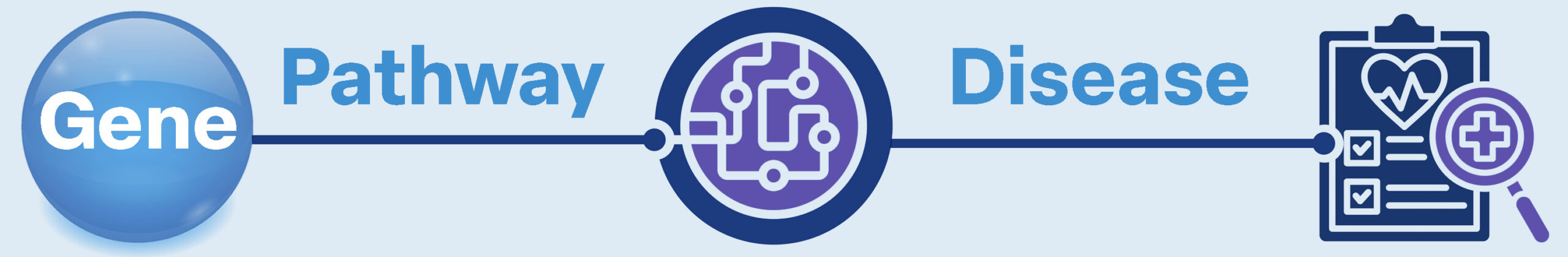
BiRAGAS models human biology as a directional causal system, connecting inherited genetic variation to molecular activity and ultimately to clinical disease.
At the foundation of the framework are genetic variants, which act as immutable causal anchors. Because genetic variation precedes all downstream biological processes, it establishes the direction of inference across regulatory mechanisms, biological programs, and clinical outcomes.
BiRAGAS framework organizes biological information into four interconnected layers.

Genetic variants (SNPs) represent fixed biological inputs that anchor the system and define the upstream origin of biological effects.

Gene expression and regulatory networks shaped by genetic variation and upstream molecular signals.

Coordinated molecular systems including pathways, biological processes, and cell-type programs that govern cellular and tissue function.

Observable phenotypes and disease states, representing the terminal layer of the causal model.
Integrated Multi-Modal Evidence
BiRAGAS integrates four complementary biological data streams into a unified causal graph.
Each pillar contributes a distinct type of evidence, and it is the convergence of all four — not any single dataset — that enables high-confidence causal inference.
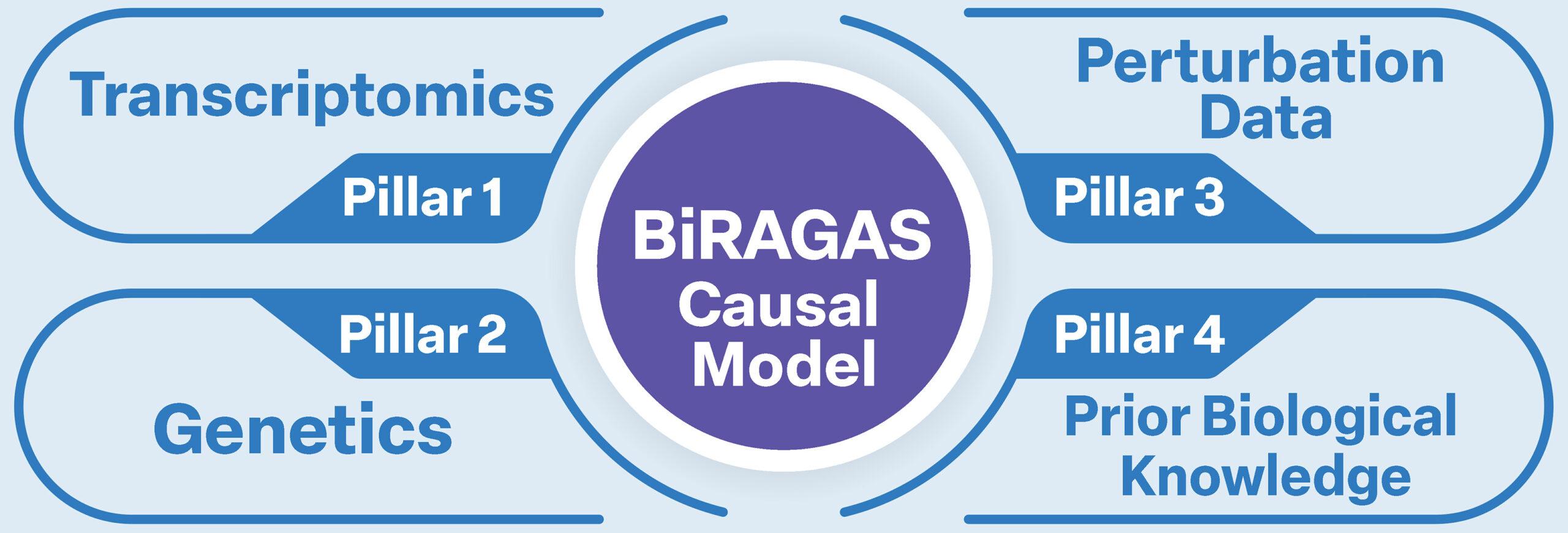

Transcriptomics
Patient cohort expression profiles across disease states.
Captures regulatory activity and differential gene expression patterns associated with disease.
Genetics
GWAS signals, eQTLs, and Mendelian randomization estimates.
Genetic variants provide the strongest causal anchor because they influence gene expression but are not altered by downstream biological events.
Perturbation Data
CRISPR DepMap essentiality screens and chemical perturbation assays. Directly tests how biological systems respond when a proposed causal gene or pathway is disrupted.
Prior Biological Knowledge
Curated pathway and interaction databases including SIGNOR, Reactome, KEGG, and Gene Ontology.
Provides biological context that validates and constrains inferred relationships.
Every Causal Claim Is Independently Tested
In BiRAGAS, no causal relationship is accepted based on a single analytical method.
Each proposed relationship is evaluated across four independent validation streams before being assigned a Causal Confidence Score.
This framework operationalizes key principles of causal inference – including strength, consistency, temporality, biological plausibility, and experimental evidence – into a rigorous computational system.

Algorithms such as PC, NOTEARS, and FCI identify candidate causal structures from large-scale observational data.

Directional time-based analyses, including Granger causality, verify that proposed causes precede their downstream effects.

Genetic instruments and eQTL signals provide natural randomization, offering strong causal evidence linking molecular changes to biological outcomes.

CRISPR perturbation screens and drug response data directly test causal hypotheses by disrupting or modulating the proposed driver.

Tiered Classification of Every Discovery
Every gene and inferred relationship in BiRAGAS is assigned a Causal Confidence Score (CCS).
This score reflects the convergence of evidence across independent validation streams and places each finding into one of three tiers.
The classification framework ensures that discoveries are prioritized according to the strength of causal evidence, guiding how results are interpreted and translated into development decisions.
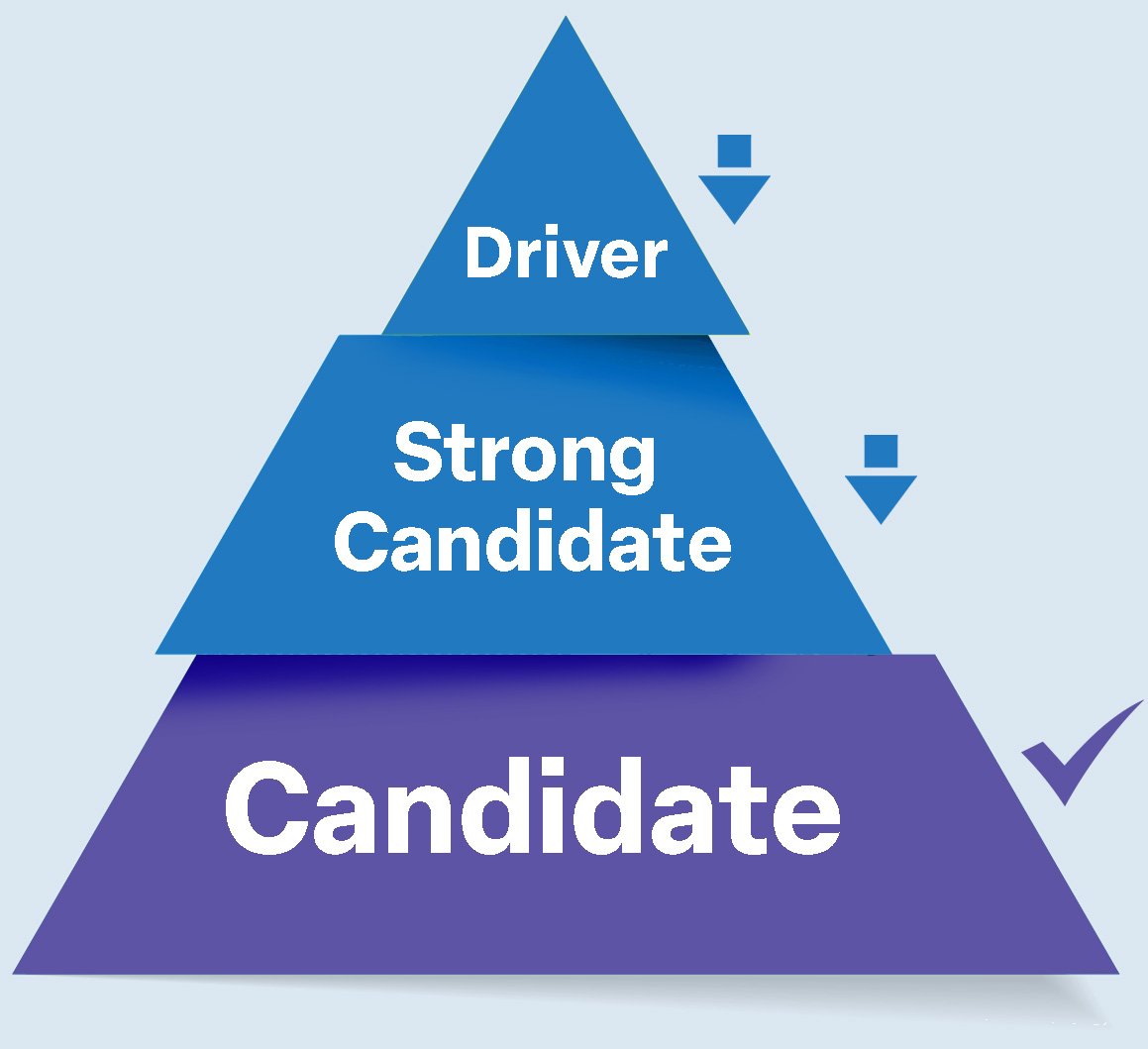
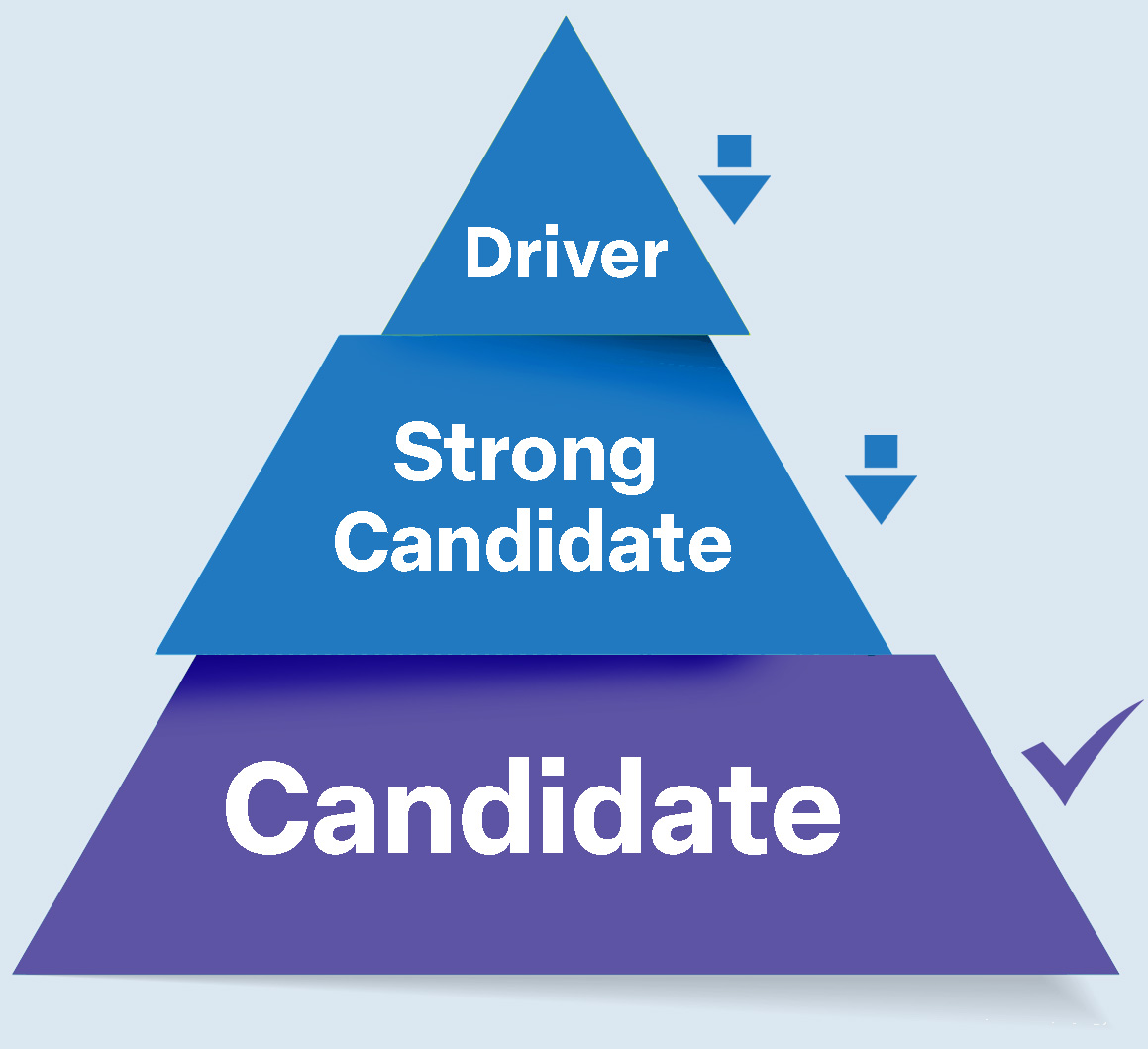
Tier 1 Driver
High-confidence causal genes supported by converging evidence across multiple independent modalities.
Appropriate candidates for direct therapeutic targeting.
Tier 2 Strong Candidate
Supported by evidence from two or more data streams.
Promising targets that warrant additional validation before full development commitment.
Tier 3 Candidate
Supported by single-modality evidence or lower confidence signals. Suitable for hypothesis generation and prioritized follow-up studies.
From Population Evidence to Individual Application
The Patient Specificity Index (PSI) framework ensures that population-level discoveries are never directly applied to individual patients without appropriate conditioning.
Each finding generated by BiRAGAS is explicitly classified by its level of patient specificity, indicating whether the evidence reflects population trends, subgroup patterns, or individual-level effects.
N ≥ 50 · Average Treatment Effect (ATE) · PSI = 0.0
Findings represent population-level averages across large patient cohorts.
These results provide broad biological insight but are not intended for direct individual prediction.
Level 2 · Subgroup
10 ≤ N < 50 · Conditional ATE · PSI = 0–1.0
Evidence applies to defined patient subgroups sharing specific biological or clinical characteristics.
These findings begin to capture conditional responses within more homogeneous populations.
Level 3 · Individual
N < 10 · Individual Treatment Effect (ITE) · PSI = 1.0
Evidence supports patient-specific inference, where causal relationships are conditioned on an individual’s molecular profile.
