BiRAGAS Framework
Autonomous Multi-Layer System for Biomarker Discovery and Transcriptomic Insight Generation
BiRAGAS bridges the gap between raw transcriptomic data and interpretable biological intelligence. Through a unified chain of RAG-driven reasoning, harmonization, differential expression analysis, cellular deconvolution, and gene prioritization, the system transforms unstructured omics data into clinically relevant insights—autonomously and reproducibly.
Built upon agentic orchestration and real-time reasoning layers, BiRAGAS reduces analytical variability by over 35% while enhancing reliability and interpretability by more than 80% across experimental contexts. The platform enables researchers and clinicians to visualize, interrogate, and validate complex molecular mechanisms within minutes – not months – thereby redefining the discovery cycle across precision medicine, immunology, and oncology.
BiRAGAS – Accelerating the future of biomarker intelligence.
BiRAGAS Causal Query Framework
From Observation Transcriptomics Query Framework
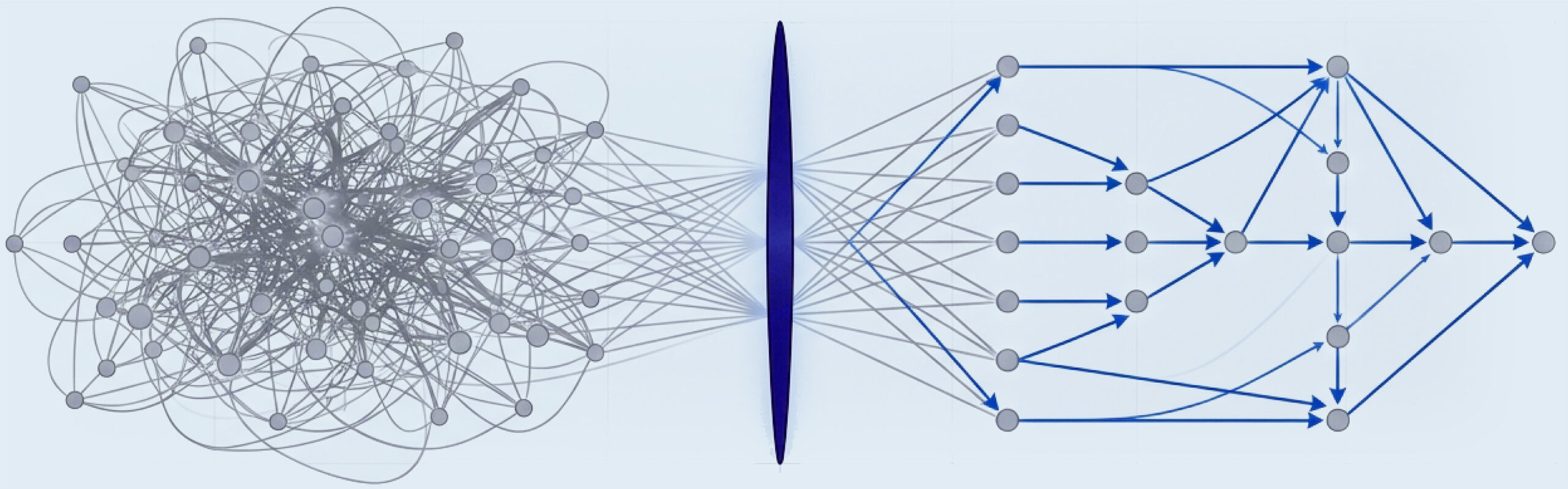
Bridging the Gap Between Observational Data and Therapeutic Intervention
The Problem
Correlation ≠ Causation
Traditional bioinformatics answers “What is different?” (association). This leaves a critical gap in drug development – because correlation is not causation. We cannot drug a consequence.
The Solution
The BiRAGAS Engine

BiRAGAS answers “What causes the difference?” It is a comprehensive engine that integrates constraint-based, score-based, and hybrid methods to address seven distinct analytical intents.
The Outcome
IND / NDA-Ready Evidence

A shift from hypothesis-generating exploration to regulatory-ready evidence.
The framework delivers de-risked targets, quantified intervention effects, and simulated counterfactuals.
A unified architecture for seven distinct models of inquiry
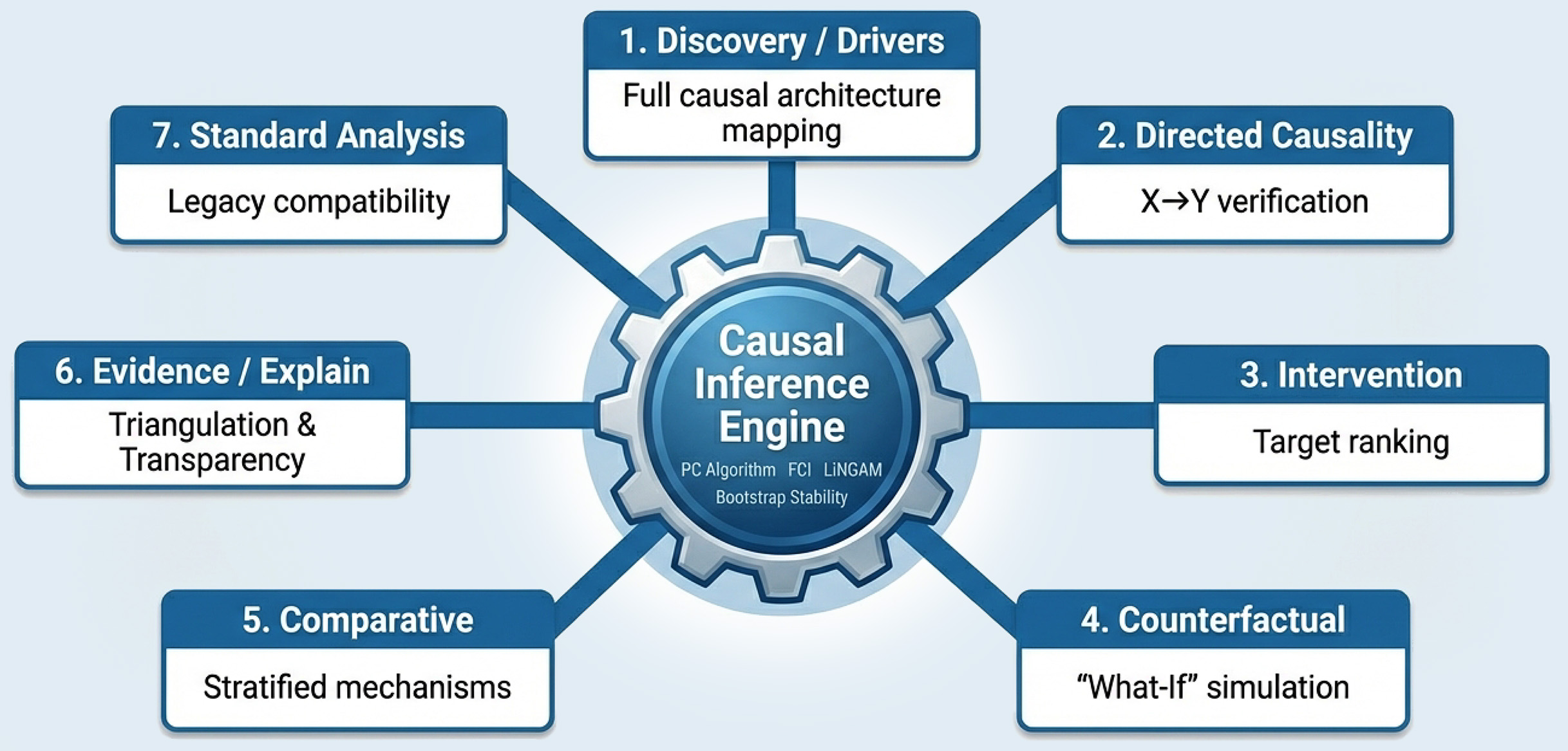
Query 1: Causal Drivers / Discovery
User Goal: Identify causal drivers of disease phenotypes through autonomous discovery
Technical Approach
-
Constraint-based: PC algorithm, FCI (latent confounders), RFCI
-
Score-based: GES (Greedy Equivalence Search)
-
Validation: Stability selection (1,000+ bootstrap iterations)
Typical Outputs
-
Ranked Gene List (Sorted by total causal effect)
-
Validated Causal Graph (Core network with >80% support)
-
Causal Equivalence Classes (Markov equivalence)
Case Study
NASH Progression
- Identified PNPLA3, TM6SF2, and MBOAT7 as top genetic drivers.
- Mapped IL-6 / TNF-α as inflammatory drivers distinct from lipotoxicity.
Query 2: Directed Causality (X → Y)
User Goal: Test whether Gene X causally affects Phenotype Y with directional specificity
Technical Approach
-
Independence Testing: Conditional independence (X ⟂ Y | Z)
-
Causal Logic: Instrumental variables (IV), do-calculus P(Y | do(X))
-
Models: LiNGAM, additive noise models (ANM)
Typical Outputs
Edge Evidence Card
-
Probability X → Y: High
-
ACE (Average Causal Effect): 0.67
-
E-Value: 4.
Case Study
Alzheimer’s Disease
- Identified PNPLA3, TM6SF2, and MBOAT7 as top genetic drivers.
- Mapped IL-6 / TNF-α as inflammatory drivers distinct from lipotoxicity.
Query 3: Intervention / Actionability
User Goal: Rank targets that effectively improve clinical outcomes if modulated
Technical Approach
-
Calculus: E[Outcome | do(Gene = intervention)]
-
Optimization: Multi-objective scoring (max effect vs. min toxicity)
-
Integration: DrugBank / ChEMBL druggability data
Typical Outputs
| Rank | Target | Druggability (0–1) | Predicted Clinical Effect |
|---|---|---|---|
| 1 | Target A | 0.95 | High |
| 2 | Target B | 0.82 | Medium |
Case Study
Diabetic Kidney Disease
- Target #1: SGLT2 (druggability: 0.95). Predicted effect: −0.8 eGFR decline.
- Target #2: JAK1/2 identified as a repurposing opportunity.
Query 4: Counterfactual What-If
User Goal: Rank targets that effectively improve clinical outcomes if modulated
Technical Approach
-
Structural Causal Models (SCM): Learning fᵧ = f(X, Uᵧ)
-
Generative AI: VAE / GAN conditioned on interventions
-
Logic: Network propagation
Typical Outputs
Shift
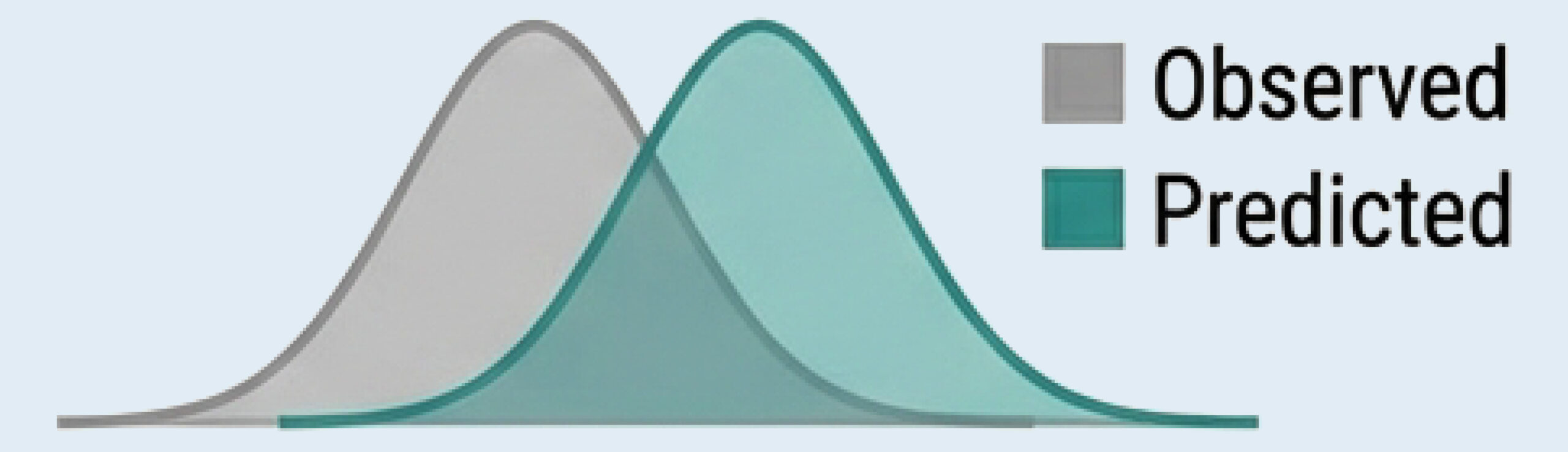
Predicted Deltas (transcriptome-wide log₂ fold change)
Uncertainty Quantification (epistemic vs. aleatoric)
Observed vs. Predicted distributions
Case Study
Beta Cell Function
- Query: Simultaneous TXNIP knockdown and PDX1 overexpression.
- Result: Predicted restoration of insulin secretion pathways (GSEA NES: 2.1) and reduction of ER stress.
Query 5: Comparative Causality
User Goal: Compare how causal mechanisms differ across patient subgroups
Technical Approach
-
Stratification: Separate inference runs per stratum
-
Differential Network Analysis: Unique edges (A vs. B)
-
Moderated Effects: Heterogeneity testing
Typical Outputs
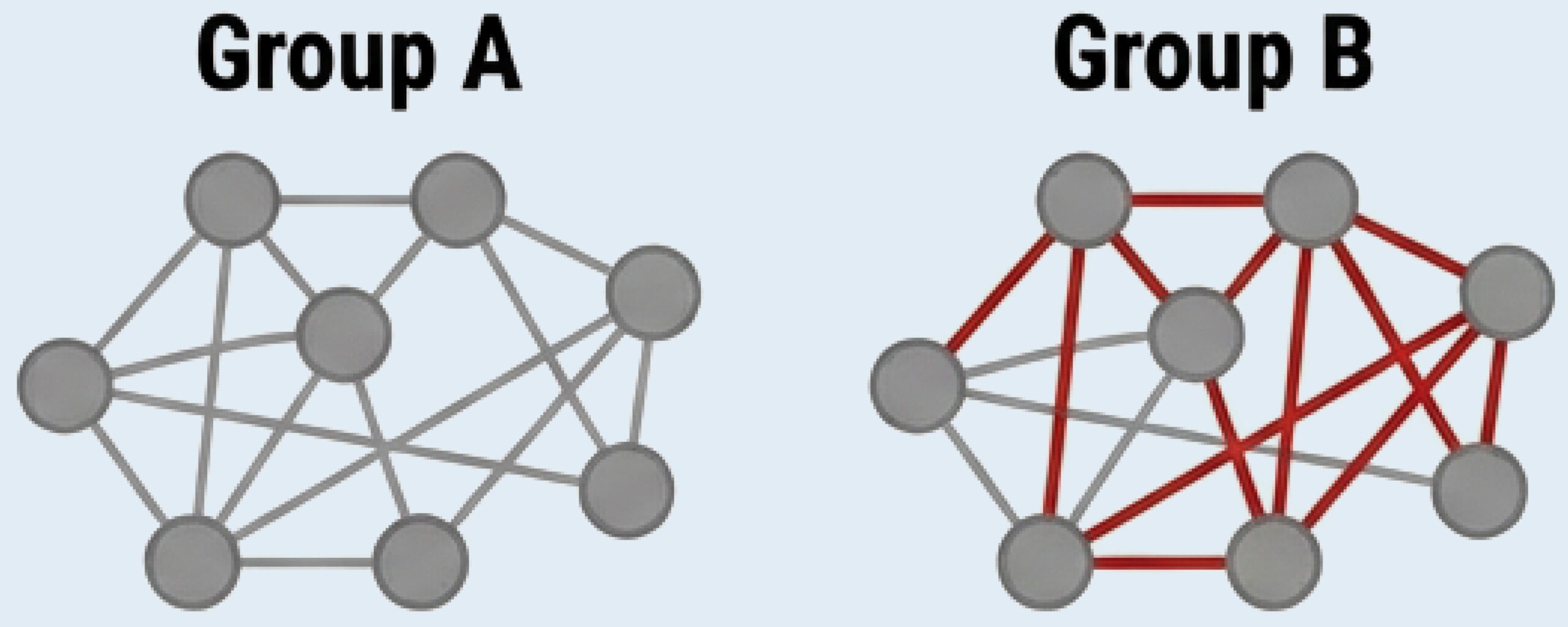
Delta Drivers Table
Case Study
Steroid Response
Responsive patients: Driven by IL-23 → Th17 axis.
Refractory patients: Driven by fibroblast-mediated ECM remodeling. Anti-inflammatories ineffective.
Query 6: Evidence Inspection / Explain
User Goal: CTransparently explain evidence, contradictions, and robustness
Technical Approach
-
Triangulation: Statistical + biological + genetic + experimental
-
Contradiction Detection: Flagging inconsistencies
-
Sensitivity: E-value calculation
Typical Outputs
Evidence Breakdown and Robustness Scores
Statistical Support: Pass
Genetic Evidence (GWAS): Pass
Literature Support: Warning, contradiction
Case Study
APOE4 Validity
Responsive patients: Driven by IL-23 → Th17 axis.
Refractory patients: Driven by fibroblast-mediated ECM remodeling. Anti-inflammatories ineffective.
Query 7: Non-Causal Standard Analysis
User Goal: CTransparently explain evidence, contradictions, and robustness
Technical Approach
-
Legacy Methods: DESeq2, limma-voom, edgeR
-
Enrichment: GSEA, ORA
-
Networks: WGCNA
Typical Outputs
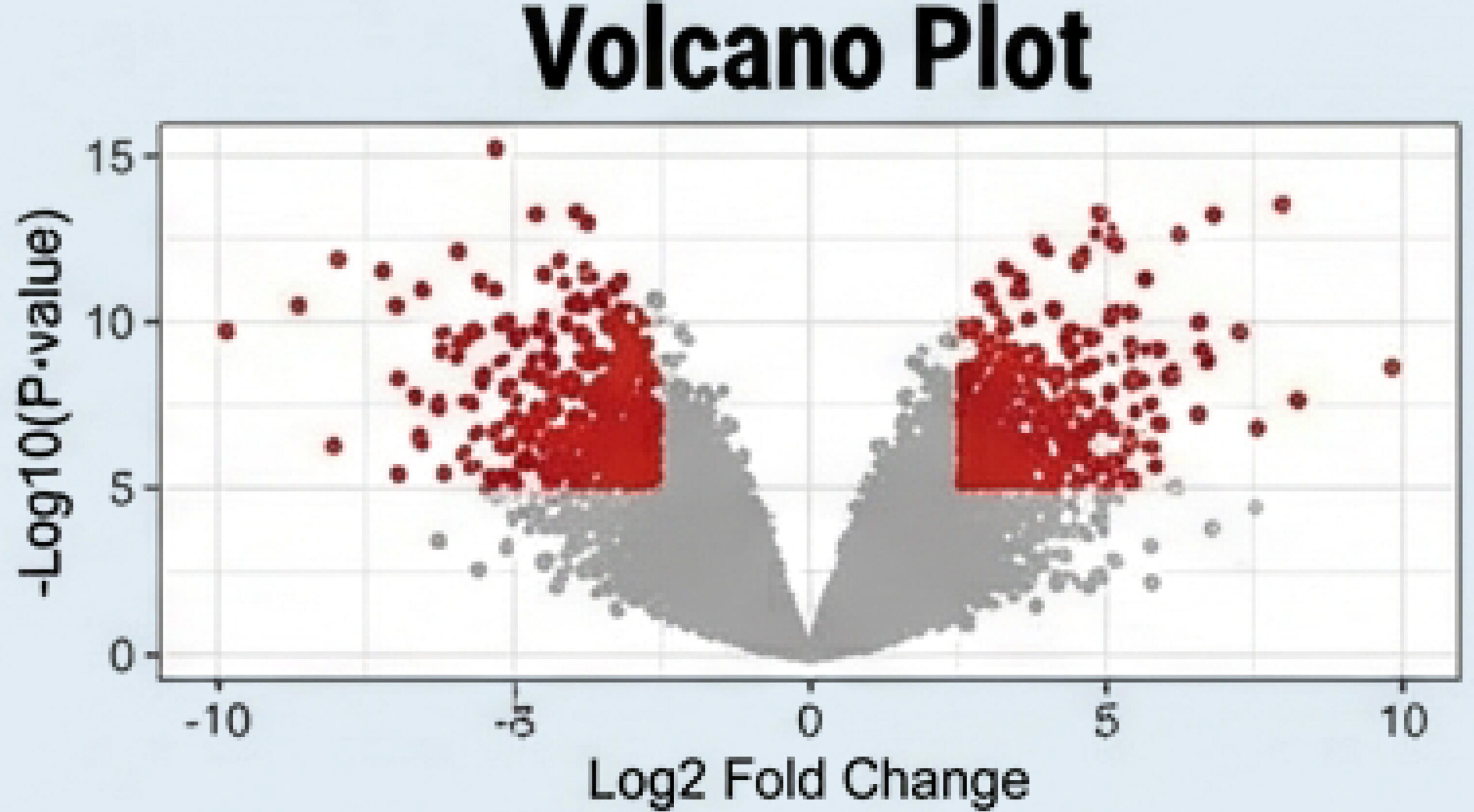
DEG Tables (log2 fold change, FDR)
Pathway Enrichment Heatmaps
Case Study
Tumor vs. Normal Colon
Standard differential expression analysis.
3,247 DEGs identified.
Top pathways include WNT signaling and cell cycle.
Integrated Worklow: From Discovery to Stratified Intervention
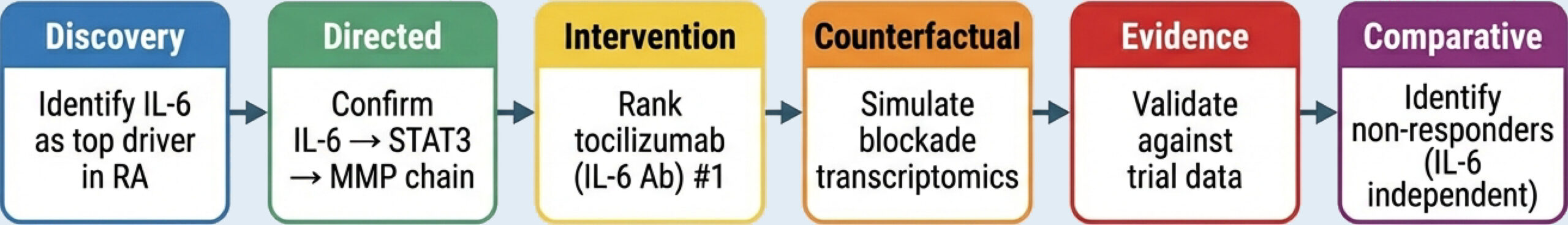
BiRAGAS supports the entire lifecycle of a therapeutic target.
Technical Differentiation
| Feature | Traditional Bioinformatics | BiRAGAS Causal Framework |
|---|---|---|
| Question Type | What is different? | What causes the difference? |
| Output | Associations / Correlations | Causal Mechanisms / Targets |
| Actionability | Hypothesis Generating | Directly Actionable / Regulatory Ready |
| Counterfactuals | Not Possible | Simulation of unobserved interventions |
| Regulatory | Exploratory | Evidence for IND/NDA |
Engineered for Robustness and Validation
Bootstrap Stability
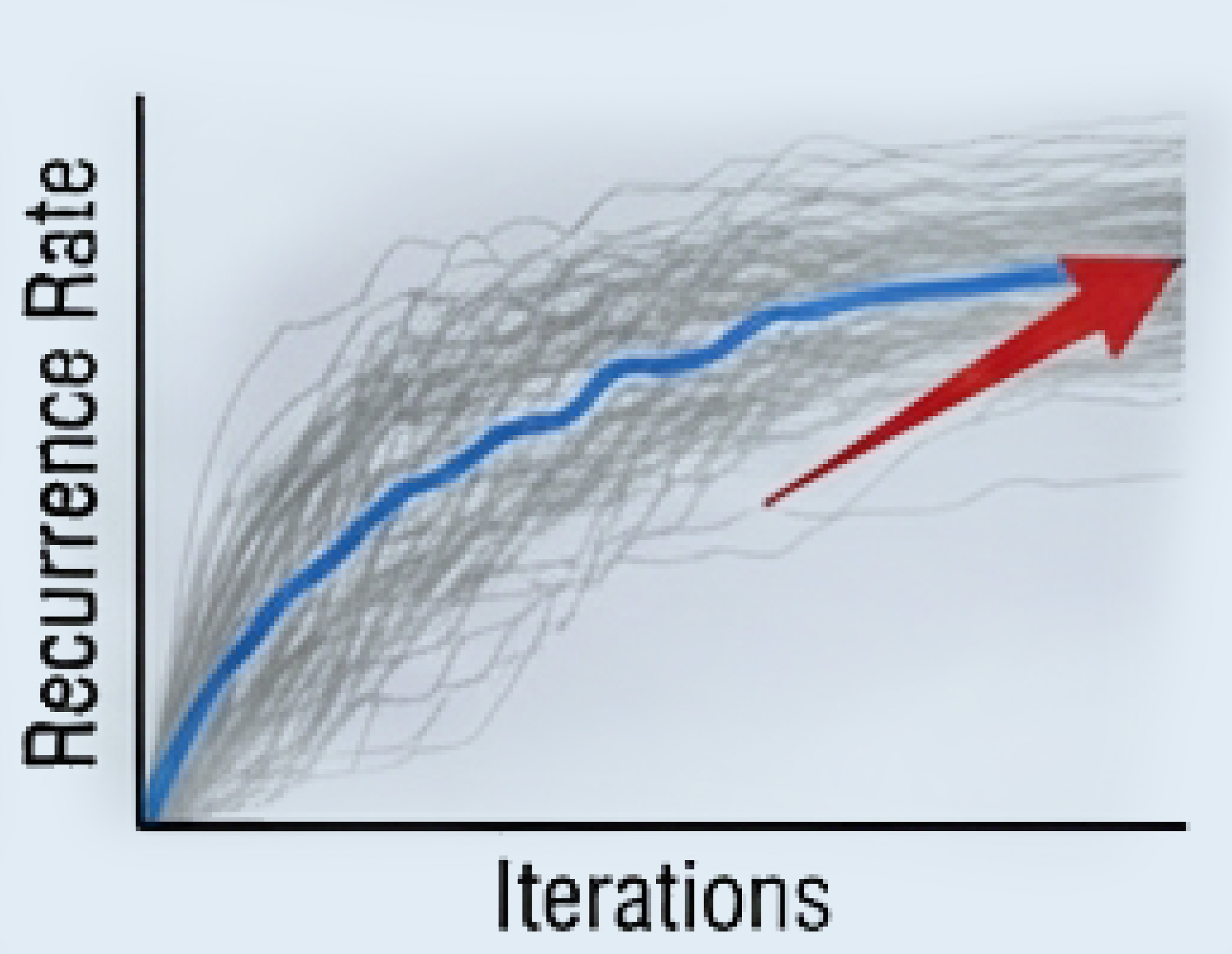
Relationships require >80% recurrence across 1000+ iterations to enter the Core Network.
Uncertainty Modeling
Explicit quantification of Epistemic (model) vs. Aleatoric (biological) uncertainty.
E-Values
E-value = RR + √[RR (RR − 1)]
Mathematical proof of required confounder strength to invalidate results.
Contradiction Analysis

Automatic downweighting of edges where biological, genetic, and statistical evidence conflict.
The BiRAGAS Competitive Edge
By systematically addressing seven distinct causal intents – from discovery to intervention to counterfactual simulation – BiRAGAS transforms transcriptomic data from descriptive associations into a competitive asset.
De-risk targets early in Discovery.
Simulate trials “in silico” before capital expenditure.
Provide transparent “White Box” evidence for regulatory bodies.
Every video featured on our website has been captured in high definition. If your device doesn’t automatically play videos in HD, kindly select the HD video quality option.
Built Ayass BioScience, LLC. Ayass BioScience, LLC If you have any questions about Ayass BioScience, LLC BiRAGAS Framework, please call today at 972-668-6005 or fill out our contact form on the bottom of this page. We will answer any question you might have.


